这个 R 包厉害了!可以在 R 语言构建多种神经网络模型,连图像分割这种复杂任务也可以完成(Rtorch教程示例)
生信碱移Rtorch经典的统计方法和简单的回归模型在面对复杂的非线性问题时,往往难以有效捕捉数据中的深层次特征。深度学习能够从数据中自动学习特征并进行复杂模式的建模,特别是在图像识别、自然语言处理和基因组学等领域表现卓越。▲ 关于神经网络的易懂讲解,可以点击此处蓝字查看小编的另一篇文章。尽管如此,生信用户最常使用的代码框架是 R 语言,而当前的主流神经网络框架 (如PyTorch) 大多基于 Py
生信碱移
Rtorch
Rtorch,顾名思义,在 R 语言中调用神经网络构建框架 Pytorch
经典的统计方法和简单的回归模型在面对复杂的非线性问题时,往往难以有效捕捉数据中的深层次特征。深度学习能够从数据中自动学习特征并进行复杂模式的建模,特别是在图像识别、自然语言处理和基因组学等领域表现卓越。
▲ 关于神经网络的易懂讲解,可以点击此处蓝字查看小编的另一篇文章。
尽管如此,生信用户最常使用的代码框架是 R 语言,而当前的主流神经网络框架 (如PyTorch) 大多基于 Python,这难免会存在一些学习成本。
Rtorch 包针对这种情况而被设计,它允许用户在 R 环境中享受PyTorch的高效梯度计算和灵活性,并处理大规模数据和进行更加个性化的模型框架设计。github链接如下:
-
https://github.com/mlverse/torch
0.推荐学习路线
① 入门教程:理解 torch 的基本使用流程和神经网络训练:
-
https://torch.mlverse.org/start/guess_the_correlation/
② 进行实验和模型调整,加深对深度学习的理解:
-
https://torch.mlverse.org/start/what_if
③ 学习创建自定义数据集,掌握数据处理技巧:
-
https://torch.mlverse.org/start/custom_dataset
④ 了解 torch 的基础构建模块,如张量、自动微分等:
-
https://torch.mlverse.org/technical/
⑤ 实际应用案例,Rtorch 在不同任务中的应用:
-
https://torch.mlverse.org/use_torch
1.R包安装与简要示例
通过Github或者CRAN皆可进行安装:
# 安装方式1
remotes::install_github("mlverse/torch")
# 安装方式2
install.packages("torch")
安装详情指引 (必看):
-
https://torch.mlverse.org/docs/articles/installation.html
可以使用torch_tensor函数从 R 对象创建 torch 张量,并使用 as_array 将它们转换回 R 对象:
library(torch)
x <- array(runif(8), dim = c(2, 2, 2))
y <- torch_tensor(x, dtype = torch_float64())
y
#> torch_tensor
#> (1,.,.) =
#> 0.6192 0.5800
#> 0.2488 0.3681
#>
#> (2,.,.) =
#> 0.0042 0.9206
#> 0.4388 0.5664
#> [ CPUDoubleType{2,2,2} ]
identical(x, as_array(y))
#> [1] TRUE
简单的自动微分示例:
x <- torch_tensor(1, requires_grad = TRUE)
w <- torch_tensor(2, requires_grad = TRUE)
b <- torch_tensor(3, requires_grad = TRUE)
y <- w * x + b
y$backward()
x$grad
#> torch_tensor
#> 2
#> [ CPUFloatType{1} ]
w$grad
#> torch_tensor
#> 1
#> [ CPUFloatType{1} ]
b$grad
#> torch_tensor
#> 1
#> [ CPUFloatType{1} ]
2.使用Rtorch进行脑图像分割
在医学中,我们可能想区分不同的细胞类型,或识别肿瘤。图像分割是一种监督学习形式:需要图片位置的真实标签。标签以掩码的形式出现,即一幅与输入数据具有相同空间分辨率的图像,标识每个像素的真实类别(此处是否为肿瘤)。因此,分类损失是逐像素计算的,然后将损失相加以得出用于优化的总和。
图像分割的常见架构是2015年发布的 U-Net。在这种架构中,存在许多变体。比如可以使用不同的层大小、激活函数、实现缩小和放大的方式等等。然而,有一个决定性的特征:U 形状,即在所有隐藏层水平交叉的连接桥。
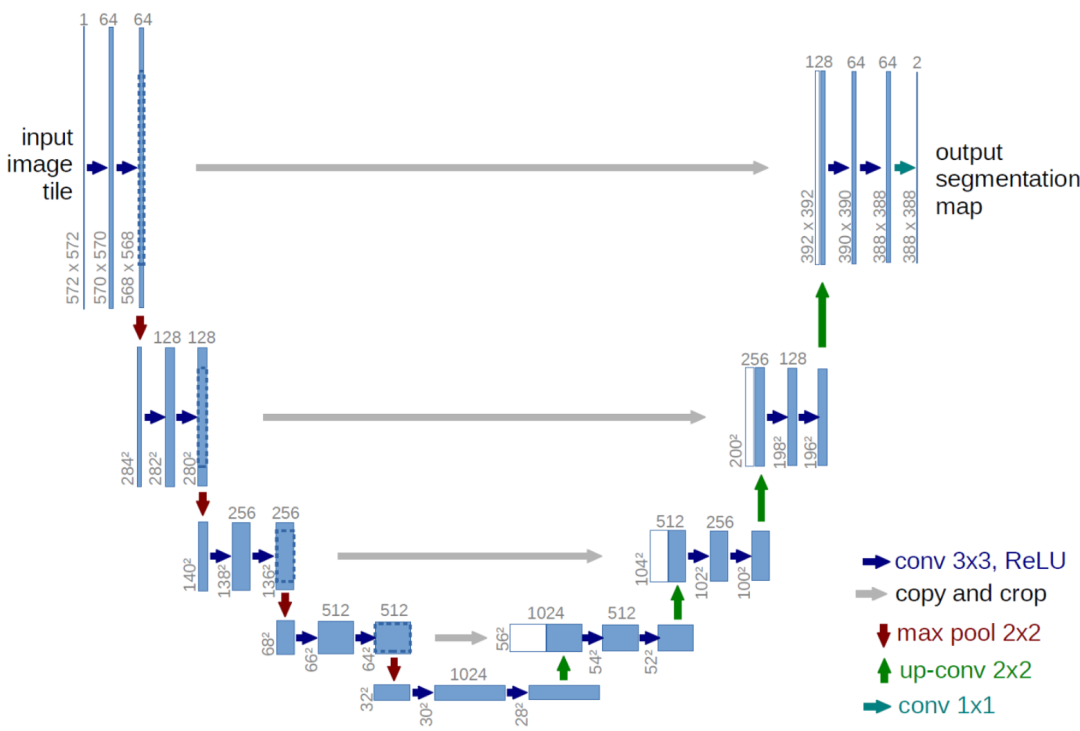
图:Unet模型架构示意图。U 的左侧类似于用于图像分类的卷积架构,逐步降低空间分辨率。然而,与分类不同,分割任务的输出应该与输入具有相同的空间分辨率。因此,需要再次放大,这由 U 的右侧处理。但是,既然降低分辨率过程中已经丢失了如此多的空间信息,模型将如何获得良好的逐像素分类?这就是中间桥的作用:在每个层级,上采样层的输入是前一层输出的拼接,该输出经过了整个压缩/解压缩过程,以及来自缩小阶段的一些保留的中间表示。通过这种方式,U-Net 架构将对细节的关注与特征提取相结合。
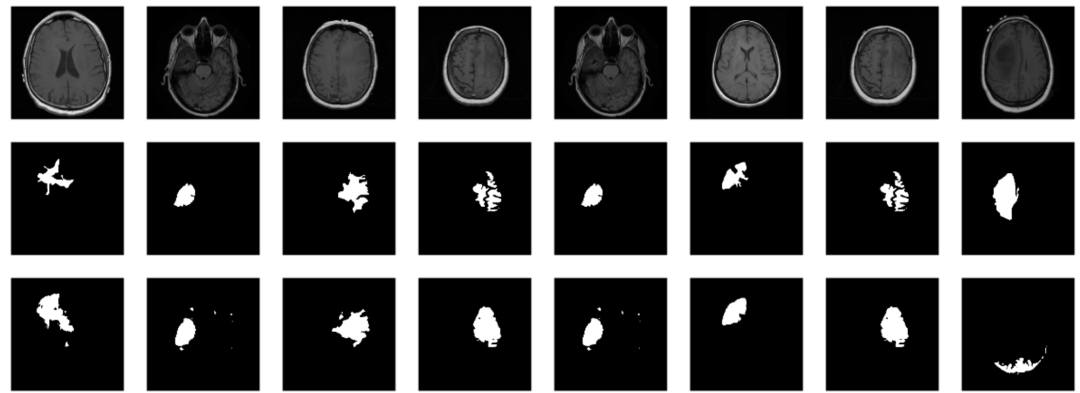
本示例演示检测脑部扫描中的异常,使用的数据集包含 MRI 图像以及手动创建的 FLAIR 异常分割掩码。即对于每位患者,多个位置都进行了切片,每位患者的切片数量不同。
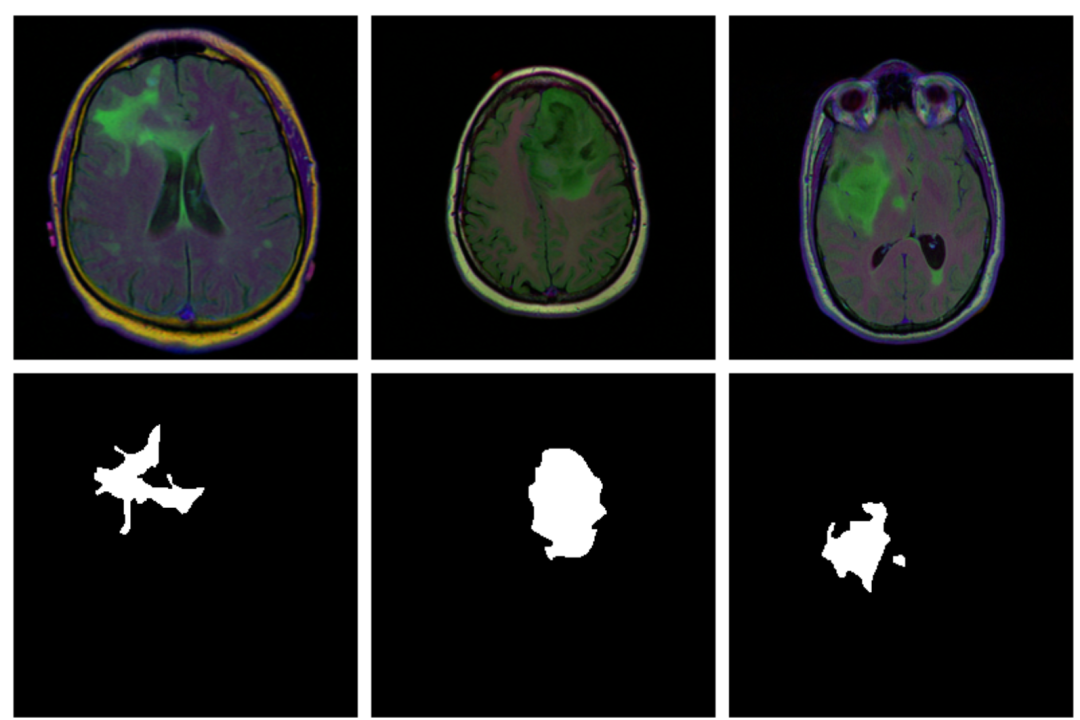
图:不同样本的脑影像(上图)以及掩码(下图)。大多数切片没有显示任何病变,相应的掩码到处都是黑色,而有病变的掩码位置标记为白色,模型的任务便是预测任意输入样本的掩码(即病变位置)。
① 使用pins包加载数据:
# deep learning (incl. dependencies)
library(torch)
library(torchvision)
# data wrangling
library(tidyverse)
library(zeallot)
# image processing and visualization
library(magick)
library(cowplot)
# dataset loading
library(pins)
library(zip)
torch_manual_seed(777)
set.seed(777)
# use your own kaggle.json here
pins::board_register_kaggle(token = "~/kaggle.json")
files <- pins::pin_get("mateuszbuda/lgg-mri-segmentation", board = "kaggle", extract = FALSE)② 创建训练与验证集:
train_dir <- "data/mri_train"
valid_dir <- "data/mri_valid"
if(dir.exists(train_dir)) unlink(train_dir, recursive = TRUE, force = TRUE)
if(dir.exists(valid_dir)) unlink(valid_dir, recursive = TRUE, force = TRUE)
zip::unzip(files, exdir = "data")
file.rename("data/kaggle_3m", train_dir)
# this is a duplicate, again containing kaggle_3m (evidently a packaging error on Kaggle)
# we just remove it
unlink("data/lgg-mri-segmentation", recursive = TRUE)
dir.create(valid_dir)
# 110 名患者中,保留 30 名用于验证
valid_indices <- sample(1:length(patients), 30)
patients <- list.dirs(train_dir, recursive = FALSE)
for (i in valid_indices) {
dir.create(file.path(valid_dir, basename(patients[i])))
for (f in list.files(patients[i])) {
file.rename(file.path(train_dir, basename(patients[i]), f), file.path(valid_dir, basename(patients[i]), f))
}
unlink(file.path(train_dir, basename(patients[i])), recursive = TRUE)
}
③ 创建torch数据加载器Dataset:
brainseg_dataset <- dataset(
name = "brainseg_dataset",
initialize = function(img_dir,
augmentation_params = NULL,
random_sampling = FALSE) {
self$images <- tibble(
img = grep(
list.files(
img_dir,
full.names = TRUE,
pattern = "tif",
recursive = TRUE
),
pattern = 'mask',
invert = TRUE,
value = TRUE
),
mask = grep(
list.files(
img_dir,
full.names = TRUE,
pattern = "tif",
recursive = TRUE
),
pattern = 'mask',
value = TRUE
)
)
self$slice_weights <- self$calc_slice_weights(self$images$mask)
self$augmentation_params <- augmentation_params
self$random_sampling <- random_sampling
},
.getitem = function(i) {
index <-
if (self$random_sampling == TRUE)
sample(1:self$.length(), 1, prob = self$slice_weights)
else
i
img <- self$images$img[index] %>%
image_read() %>%
transform_to_tensor()
mask <- self$images$mask[index] %>%
image_read() %>%
transform_to_tensor() %>%
transform_rgb_to_grayscale() %>%
torch_unsqueeze(1)
img <- self$min_max_scale(img)
if (!is.null(self$augmentation_params)) {
scale_param <- self$augmentation_params[1]
c(img, mask) %<-% self$resize(img, mask, scale_param)
rot_param <- self$augmentation_params[2]
c(img, mask) %<-% self$rotate(img, mask, rot_param)
flip_param <- self$augmentation_params[3]
c(img, mask) %<-% self$flip(img, mask, flip_param)
}
list(img = img, mask = mask)
},
.length = function() {
nrow(self$images)
},
calc_slice_weights = function(masks) {
weights <- map_dbl(masks, function(m) {
img <-
as.integer(magick::image_data(image_read(m), channels = "gray"))
sum(img / 255)
})
sum_weights <- sum(weights)
num_weights <- length(weights)
weights <- weights %>% map_dbl(function(w) {
w <- (w + sum_weights * 0.1 / num_weights) / (sum_weights * 1.1)
})
weights
},
min_max_scale = function(x) {
min = x$min()$item()
max = x$max()$item()
x$clamp_(min = min, max = max)
x$add_(-min)$div_(max - min + 1e-5)
x
},
resize = function(img, mask, scale_param) {
img_size <- dim(img)[2]
rnd_scale <- runif(1, 1 - scale_param, 1 + scale_param)
img <- transform_resize(img, size = rnd_scale * img_size)
mask <- transform_resize(mask, size = rnd_scale * img_size)
diff <- dim(img)[2] - img_size
if (diff > 0) {
top <- ceiling(diff / 2)
left <- ceiling(diff / 2)
img <- transform_crop(img, top, left, img_size, img_size)
mask <- transform_crop(mask, top, left, img_size, img_size)
} else {
img <- transform_pad(img,
padding = -c(
ceiling(diff / 2),
floor(diff / 2),
ceiling(diff / 2),
floor(diff / 2)
))
mask <- transform_pad(mask, padding = -c(
ceiling(diff / 2),
floor(diff /
2),
ceiling(diff /
2),
floor(diff /
2)
))
}
list(img, mask)
},
rotate = function(img, mask, rot_param) {
rnd_rot <- runif(1, 1 - rot_param, 1 + rot_param)
img <- transform_rotate(img, angle = rnd_rot)
mask <- transform_rotate(mask, angle = rnd_rot)
list(img, mask)
},
flip = function(img, mask, flip_param) {
rnd_flip <- runif(1)
if (rnd_flip > flip_param) {
img <- transform_hflip(img)
mask <- transform_hflip(mask)
}
list(img, mask)
}
)
train_ds <- brainseg_dataset(
train_dir,
augmentation_params = c(0.05, 15, 0.5),
random_sampling = TRUE
)
length(train_ds)
# 2977
valid_ds <- brainseg_dataset(
valid_dir,
augmentation_params = NULL,
random_sampling = FALSE
)
length(valid_ds)
# 952
④ 可视化一个样本与其掩码(已知病变标签位置):
par(mfrow = c(1, 2), mar = c(0, 1, 0, 1))
img_and_mask <- valid_ds[27]
img <- img_and_mask[[1]]
mask <- img_and_mask[[2]]
img$permute(c(2, 3, 1)) %>% as.array() %>% as.raster() %>% plot()
mask$squeeze() %>% as.array() %>% as.raster() %>% plot()
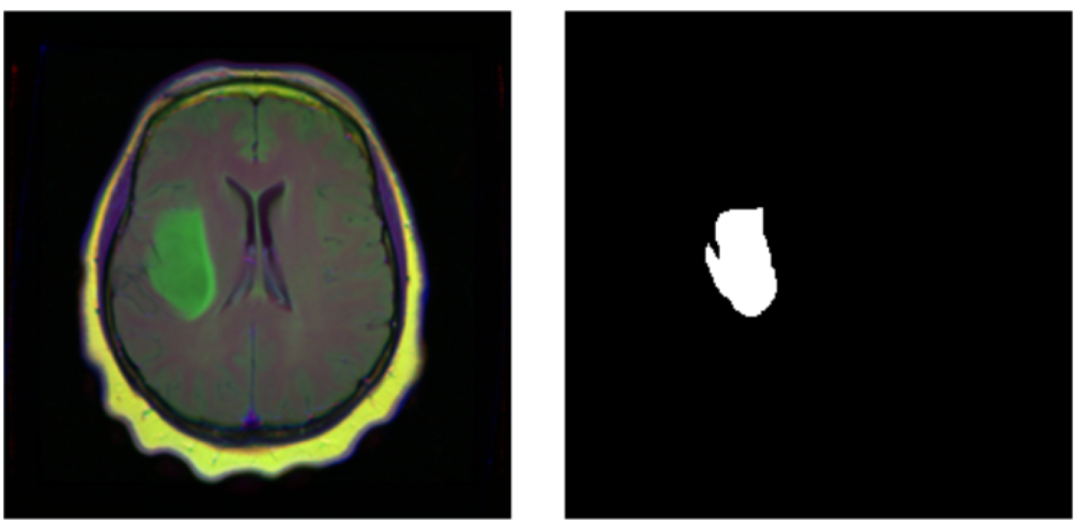
⑤ 查看不同增强参数对验证集数据的影响:
img_and_mask <- valid_ds[77]
img <- img_and_mask[[1]]
mask <- img_and_mask[[2]]
imgs <- map (1:24, function(i) {
# scale factor; train_ds really uses 0.05
c(img, mask) %<-% valid_ds$resize(img, mask, 0.2)
c(img, mask) %<-% valid_ds$flip(img, mask, 0.5)
# rotation angle; train_ds really uses 15
c(img, mask) %<-% valid_ds$rotate(img, mask, 90)
img %>%
transform_rgb_to_grayscale() %>%
as.array() %>%
as_tibble() %>%
rowid_to_column(var = "Y") %>%
gather(key = "X", value = "value", -Y) %>%
mutate(X = as.numeric(gsub("V", "", X))) %>%
ggplot(aes(X, Y, fill = value)) +
geom_raster() +
theme_void() +
theme(legend.position = "none") +
theme(aspect.ratio = 1)
})
plot_grid(plotlist = imgs, nrow = 4)
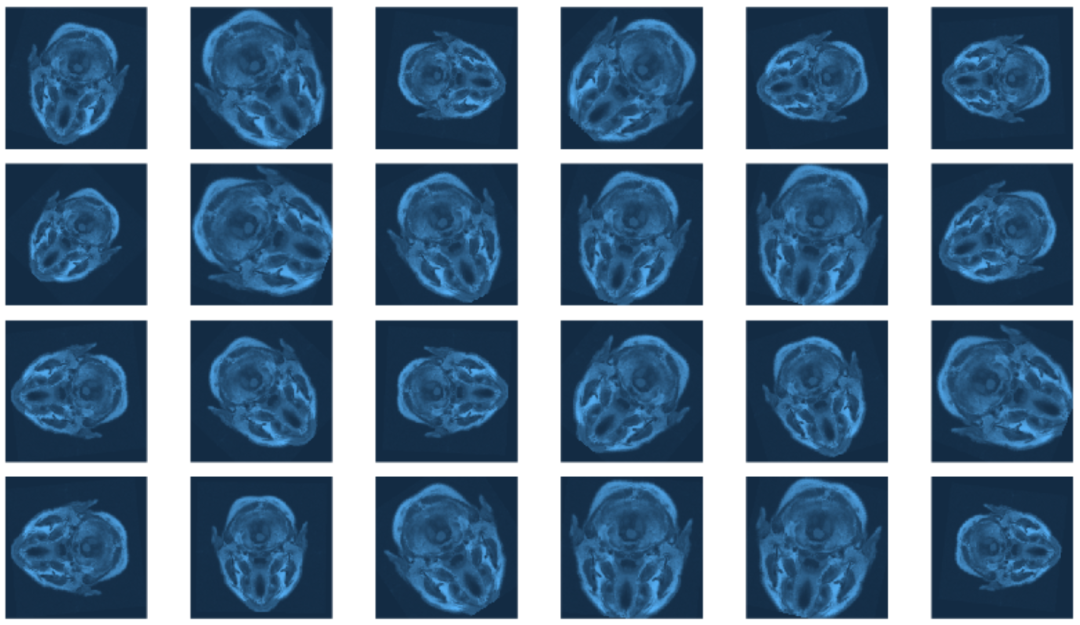

⑥ 创建数据加载器,设置合适的批次,越大的迭代将会更快但是更加消耗资源:
batch_size <- 4 # 一次加载4个样本
train_dl <- dataloader(train_ds, batch_size)
valid_dl <- dataloader(valid_ds, batch_size)
⑦ 模型架构搭建:
unet <- nn_module(
"unet",
initialize = function(channels_in = 3,
n_classes = 1,
depth = 5,
n_filters = 6) {
self$down_path <- nn_module_list()
prev_channels <- channels_in
for (i in 1:depth) {
self$down_path$append(down_block(prev_channels, 2 ^ (n_filters + i - 1)))
prev_channels <- 2 ^ (n_filters + i -1)
}
self$up_path <- nn_module_list()
for (i in ((depth - 1):1)) {
self$up_path$append(up_block(prev_channels, 2 ^ (n_filters + i - 1)))
prev_channels <- 2 ^ (n_filters + i - 1)
}
self$last = nn_conv2d(prev_channels, n_classes, kernel_size = 1)
},
forward = function(x) {
blocks <- list()
for (i in 1:length(self$down_path)) {
x <- self$down_path[[i]](x)
if (i != length(self$down_path)) {
blocks <- c(blocks, x)
x <- nnf_max_pool2d(x, 2)
}
}
for (i in 1:length(self$up_path)) {
x <- self$up_path[[i]](x, blocks[[length(blocks) - i + 1]]$to(device = device))
}
torch_sigmoid(self$last(x))
}
)
down_block <- nn_module(
"down_block",
initialize = function(in_size, out_size) {
self$conv_block <- conv_block(in_size, out_size)
},
forward = function(x) {
self$conv_block(x)
}
)
up_block <- nn_module(
"up_block",
initialize = function(in_size, out_size) {
self$up = nn_conv_transpose2d(in_size,
out_size,
kernel_size = 2,
stride = 2)
self$conv_block = conv_block(in_size, out_size)
},
forward = function(x, bridge) {
up <- self$up(x)
torch_cat(list(up, bridge), 2) %>%
self$conv_block()
}
)
conv_block <- nn_module(
"conv_block",
initialize = function(in_size, out_size) {
self$conv_block <- nn_sequential(
nn_conv2d(in_size, out_size, kernel_size = 3, padding = 1),
nn_relu(),
nn_dropout(0.6),
nn_conv2d(out_size, out_size, kernel_size = 3, padding = 1),
nn_relu()
)
},
forward = function(x){
self$conv_block(x)
}
)
# 实例化
device <- torch_device(if(cuda_is_available()) "cuda" else "cpu")
model <- unet(depth = 5)$to(device = device)
⑧ 设计损失函数以及模型训练:
calc_dice_loss <- function(y_pred, y_true) {
smooth <- 1
y_pred <- y_pred$view(-1)
y_true <- y_true$view(-1)
intersection <- (y_pred * y_true)$sum()
1 - ((2 * intersection + smooth) / (y_pred$sum() + y_true$sum() + smooth))
}
dice_weight <- 0.3
optimizer <- optim_sgd(model$parameters, lr = 0.1, momentum = 0.9)
num_epochs <- 20
scheduler <- lr_one_cycle(
optimizer,
max_lr = 0.1,
steps_per_epoch = length(train_dl),
epochs = num_epochs
)
train_batch <- function(b) {
optimizer$zero_grad()
output <- model(b[[1]]$to(device = device))
target <- b[[2]]$to(device = device)
bce_loss <- nnf_binary_cross_entropy(output, target)
dice_loss <- calc_dice_loss(output, target)
loss <- dice_weight * dice_loss + (1 - dice_weight) * bce_loss
loss$backward()
optimizer$step()
scheduler$step()
list(bce_loss$item(), dice_loss$item(), loss$item())
}
valid_batch <- function(b) {
output <- model(b[[1]]$to(device = device))
target <- b[[2]]$to(device = device)
bce_loss <- nnf_binary_cross_entropy(output, target)
dice_loss <- calc_dice_loss(output, target)
loss <- dice_weight * dice_loss + (1 - dice_weight) * bce_loss
list(bce_loss$item(), dice_loss$item(), loss$item())
}
for (epoch in 1:num_epochs) {
model$train()
train_bce <- c()
train_dice <- c()
train_loss <- c()
coro::loop(for (b in train_dl) {
c(bce_loss, dice_loss, loss) %<-% train_batch(b)
train_bce <- c(train_bce, bce_loss)
train_dice <- c(train_dice, dice_loss)
train_loss <- c(train_loss, loss)
})
torch_save(model, paste0("model_", epoch, ".pt"))
cat(sprintf("\nEpoch %d, training: loss:%3f, bce: %3f, dice: %3f\n",
epoch, mean(train_loss), mean(train_bce), mean(train_dice)))
model$eval()
valid_bce <- c()
valid_dice <- c()
valid_loss <- c()
i <- 0
coro::loop(for (b in tvalid_dl) {
i <<- i + 1
c(bce_loss, dice_loss, loss) %<-% valid_batch(b)
valid_bce <- c(valid_bce, bce_loss)
valid_dice <- c(valid_dice, dice_loss)
valid_loss <- c(valid_loss, loss)
})
cat(sprintf("\nEpoch %d, validation: loss:%3f, bce: %3f, dice: %3f\n",
epoch, mean(valid_loss), mean(valid_bce), mean(valid_dice)))
}
#Epoch 1, training: loss:0.304232, bce: 0.148578, dice: 0.667423
#Epoch 1, validation: loss:0.333961, bce: 0.127171, dice: 0.816471
#Epoch 2, training: loss:0.194665, bce: 0.101973, dice: 0.410945
#Epoch 2, validation: loss:0.341121, bce: 0.117465, dice: 0.862983
#[...]
#Epoch 19, training: loss:0.073863, bce: 0.038559, dice: 0.156236
#Epoch 19, validation: loss:0.302878, bce: 0.109721, dice: 0.753577
#Epoch 20, training: loss:0.070621, bce: 0.036578, dice: 0.150055
#Epoch 20, validation: loss:0.295852, bce: 0.101750, dice: 0.748757
⑨ 模型评估与预测,对未知的样本预测其病变位置:
saved_model <- torch_load("model_20.pt")
model <- saved_model
model$eval()
# without random sampling, we'd mainly see lesion-free patches
eval_ds <- brainseg_dataset(valid_dir, augmentation_params = NULL, random_sampling = TRUE)
eval_dl <- dataloader(eval_ds, batch_size = 8)
batch <- eval_dl %>% dataloader_make_iter() %>% dataloader_next()
par(mfcol = c(3, 8), mar = c(0, 1, 0, 1))
for (i in 1:8) {
img <- batch[[1]][i, .., drop = FALSE]
inferred_mask <- model(img$to(device = device))
true_mask <- batch[[2]][i, .., drop = FALSE]$to(device = device)
bce <- nnf_binary_cross_entropy(inferred_mask, true_mask)$to(device = "cpu") %>%
as.numeric()
dc <- calc_dice_loss(inferred_mask, true_mask)$to(device = "cpu") %>% as.numeric()
cat(sprintf("\nSample %d, bce: %3f, dice: %3f\n", i, bce, dc))
inferred_mask <- inferred_mask$to(device = "cpu") %>% as.array() %>% .[1, 1, , ]
inferred_mask <- ifelse(inferred_mask > 0.5, 1, 0)
img[1, 1, ,] %>% as.array() %>% as.raster() %>% plot()
true_mask$to(device = "cpu")[1, 1, ,] %>% as.array() %>% as.raster() %>% plot()
inferred_mask %>% as.raster() %>% plot()
}
感觉确实灵活?
大家可以使用试试

魔乐社区(Modelers.cn) 是一个中立、公益的人工智能社区,提供人工智能工具、模型、数据的托管、展示与应用协同服务,为人工智能开发及爱好者搭建开放的学习交流平台。社区通过理事会方式运作,由全产业链共同建设、共同运营、共同享有,推动国产AI生态繁荣发展。
更多推荐
 已为社区贡献11条内容
已为社区贡献11条内容






所有评论(0)