transunet训练自己的数据集,测试数据值太低
用WORD数据集做腹部多器官分割任务时,将数据集转换成相应的.npz格式和.npy.h5格式,测试数据如图,测试数据中的 mean_dice 和 mean_hd95 值太低,是由于什么原因造成的?3.在官方代码下的问答发现是否是utils.py中对dice和hd95的计算有关?2.换数据集后dataset_synapse.py需要修改,但我没有修改?1.转换.npz和.npy.h5文件出错,转换代
·
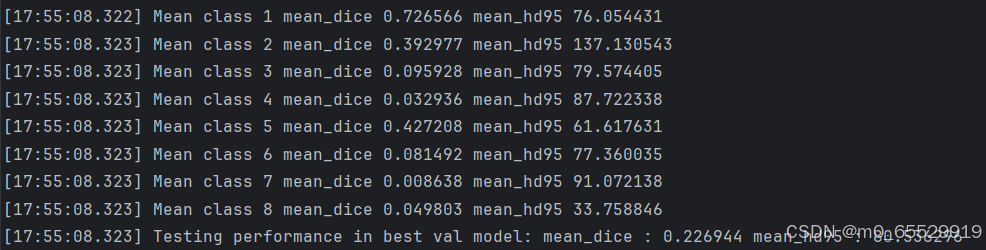
用WORD数据集做腹部多器官分割任务时,将数据集转换成相应的.npz格式和.npy.h5格式,测试数据如图,测试数据中的 mean_dice 和 mean_hd95 值太低,是由于什么原因造成的?
预测原因如下:
1.转换.npz和.npy.h5文件出错,转换代码如下
2.换数据集后dataset_synapse.py需要修改,但我没有修改?
3.在官方代码下的问答发现是否是utils.py中对dice和hd95的计算有关?
一直找不到原因在哪里,如果有大佬告诉我原因出在哪里就太好了,谢谢!!!
转换npz和npy.h5代码如下
import numpy as np
import nibabel as nib
import h5py
import os
from scipy.ndimage import zoom
# 设定路径
base_path = r"E:\Semantic Segmentation\tradition\transUnet\project_TransUNet(1)"
train_img_folder = os.path.join(base_path, r"E:\SemanticSegmentation\database\word\Training-Testing\img\imageTr")
test_img_folder = os.path.join(base_path, r"E:\SemanticSegmentation\database\word\Training-Testing\img\imageTs")
train_label_folder = os.path.join(base_path, r"E:\SemanticSegmentation\database\word\Training-Testing\label\labelTr")
test_label_folder = os.path.join(base_path, r"E:\SemanticSegmentation\database\word\Training-Testing\label\labelTs")
train_save_folder = os.path.join(base_path, r"E:\SemanticSegmentation\tradition\transUnet\project_TransUNet(1)\data\Synapse\train_npz")
test_save_folder = os.path.join(base_path, r"E:\SemanticSegmentation\tradition\transUnet\project_TransUNet(1)\data\Synapse\test_vol_h5")
# 创建保存路径
os.makedirs(train_save_folder, exist_ok=True)
os.makedirs(test_save_folder, exist_ok=True)
def filter_labels(label_data, key_labels=[1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16]):
"""
将标签中保留的关键标签设为原值,其余标签设为0
"""
filtered_labels = np.zeros_like(label_data)
for key in key_labels:
filtered_labels[label_data == key] = key
return filtered_labels
def contains_key_labels(slice_label, key_labels=[1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16]):
"""
检查切片是否包含关键标签,且背景占比不为 100%
"""
total_pixels = slice_label.size
background_pixels = np.sum(slice_label == 0)
background_ratio = background_pixels / total_pixels
contains_labels = np.isin(slice_label, key_labels).any()
# 只保留包含关键标签且背景占比不为 100% 的切片
return contains_labels or background_ratio < 1.0
# 修改后的测试集数据保存逻辑
for filename in os.listdir(test_img_folder):
if filename.endswith('.nii.gz'):
img_path = os.path.join(test_img_folder, filename)
label_path = os.path.join(test_label_folder, filename) if os.path.exists(
os.path.join(test_label_folder, filename)) else None
# 加载图像和标签
img = nib.load(img_path)
data = img.get_fdata()
label_data = nib.load(label_path).get_fdata() if label_path else np.zeros(data.shape) # 如果没有标签,填充全0
# 直接使用原始数据,无需重采样
data_resampled = data
label_resampled = label_data
# 只保留标签1-8,其他标签归为0
label_resampled = filter_labels(label_resampled)
# 查找包含关键标签的第一个和最后一个切片索引
start_slice = None
end_slice = None
for slice_idx in range(label_resampled.shape[2]):
if contains_key_labels(label_resampled[:, :, slice_idx]):
if start_slice is None:
start_slice = slice_idx
end_slice = slice_idx # 不断更新end_slice,直到最后一个关键标签切片
# 如果找到关键标签的范围
if start_slice is not None and end_slice is not None:
# 裁剪数据到包含关键标签的切片范围
data_clipped = data_resampled[:, :, start_slice:end_slice+1]
label_clipped = label_resampled[:, :, start_slice:end_slice+1]
# 数据裁剪和归一化
data_clipped = np.clip(data_clipped, -125, 275)
data_normalized = (data_clipped - (-125)) / (275 - (-125))
# 保存3D体数据为 .npy.h5 文件
num_slices = data_clipped.shape[2]
with h5py.File(os.path.join(test_save_folder, f"case_{filename.split('.')[0]}.npy.h5"), 'w') as f:
f.create_dataset("image", data=data_clipped)
f.create_dataset("label", data=label_clipped)
# 打印处理完成的信息
print(f"文件 {filename} 中处理完成,共生成 {num_slices} 张切片。")
else:
print(f"文件 {filename} 中没有发现关键标签的切片。")
# 修改后的训练集数据保存逻辑
for filename in os.listdir(train_img_folder):
if filename.endswith('.nii.gz'):
img_path = os.path.join(train_img_folder, filename)
label_path = os.path.join(train_label_folder, filename) if os.path.exists(
os.path.join(train_label_folder, filename)) else None
# 加载图像和标签
img = nib.load(img_path)
data = img.get_fdata()
label_data = nib.load(label_path).get_fdata() if label_path else np.zeros(data.shape) # 如果没有标签,填充全0
# 直接使用原始数据,无需重采样
data_resampled = data
label_resampled = filter_labels(label_data)
# 查找包含关键标签的第一个和最后一个切片索引
start_slice = None
end_slice = None
for slice_idx in range(label_resampled.shape[2]):
if contains_key_labels(label_resampled[:, :, slice_idx]):
if start_slice is None:
start_slice = slice_idx
end_slice = slice_idx # 不断更新end_slice,直到最后一个关键标签切片
# 如果找到关键标签的范围
if start_slice is not None and end_slice is not None:
# 裁剪数据到包含关键标签的切片范围
data_clipped = data_resampled[:, :, start_slice:end_slice + 1]
label_clipped = label_resampled[:, :, start_slice:end_slice + 1]
# 数据裁剪和归一化
data_clipped = np.clip(data_clipped, -125, 275)
data_normalized = (data_clipped - (-125)) / (275 - (-125))
# 保存每个切片为单独的 .npz 文件
for i in range(data_normalized.shape[2]):
slice_image = data_normalized[:, :, i]
slice_label = label_clipped[:, :, i]
np.savez_compressed(
os.path.join(train_save_folder, f"case_{filename.split('.')[0]}_slice_{i:03d}.npz"),
image=slice_image,
label=slice_label
)
# 打印处理完成的信息
print(f"文件 {filename} 处理完成并保存为多个 .npz 格式。")
else:
print(f"文件 {filename} 中没有发现关键标签的切片。")
魔乐社区(Modelers.cn) 是一个中立、公益的人工智能社区,提供人工智能工具、模型、数据的托管、展示与应用协同服务,为人工智能开发及爱好者搭建开放的学习交流平台。社区通过理事会方式运作,由全产业链共同建设、共同运营、共同享有,推动国产AI生态繁荣发展。
更多推荐
 已为社区贡献1条内容
已为社区贡献1条内容






所有评论(0)